Project Overview
We use a combination of quantitative fluorescence microscopy, super resolution imaging, and mathematical modeling to understand how the structural units of the contractile ring assemble in fission yeast cells. These structural units, which we call nodes, are discrete protein complexes sitting on the inner surface of the cell that self-assemble into the contractile ring using actin and myosin. The contractile ring constricts to split the cell in half during cytokinesis. Quantitative measurements and modeling had revealed the mechanism by which nodes assemble into the ring during mitosis, but little was known about how these nodes assembled earlier in the cell cycle, during interphase.
We found that fission yeast use two distinct types of nodes in interphase (fluorescence image) which form separately and then merge at equator of the cell to prepare nodes for cytokinesis. One type of node forms around the mother cell’s divided daughter nuclei (summary cartoon) before the cell splits in half into two daughter cells. The second type of node emerges from the disassembling contractile ring of the mother cell, which is the new tip of the two resultant daughter cells.
We tracked fluorescently labeled nodes in live cells to propose a mechanism by which both types of nodes merge at the equator of daughter cells: Type 2 nodes diffuse along the cell surface from their origin at the contractile ring until they are captured at the equator of the daughter cell by stationary Type 1 nodes. This “diffuse-and capture” mechanism was supported by stochastic simulations of a mathematical model (simulation video). The two types of nodes, united at the cell equator, accumulate cytokinesis proteins to assemble the contractile ring and then separate, thereby completing their journey around the cell cycle. The two types of nodes therefore follow parallel branches of the pathway that prepares nodes for cytokinesis. Most recently, we started using super resolution imaging to study how the cytokinetic nodes assemble in live cells.
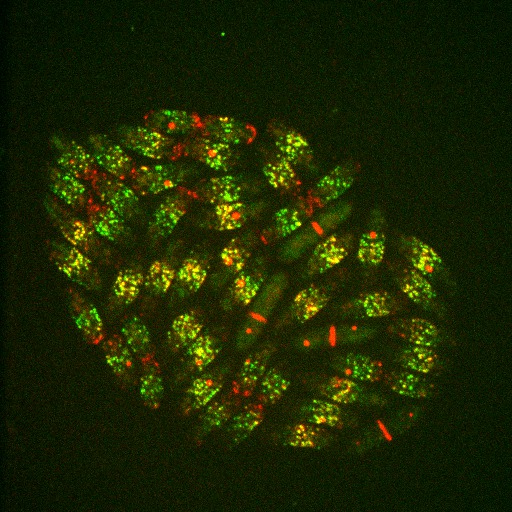
Image above: Fluorescence image of rod-shaped fission yeast cells expressing fluorescent markers for two types of nodes. Type 1 nodes (green) are separate from Type 2 nodes (red) in some cells, but in other cells the two types of nodes are located within the same fluorescent spots (yellow). During mitosis (long cells) Type 2 nodes are located within the contractile ring in the middle of the cell while the protein contents of Type 1 nodes are dispersed into the cell cytoplasm. Spindle pole bodies (red spots) were used as an internal timer for the cellular clock.
Video above: Video output of diffuse-and-capture simulation. In the stochastic simulation (made in MATLAB), Type 2 nodes (red) started at the cell tip (top) and diffused in two dimensions until they were captured (yellow) by passing near a Type 1 node (green) located near the cell equator. Time is in hours: minutes: seconds. The white box outlines the dimensions of cells projected in our fluorescence images.
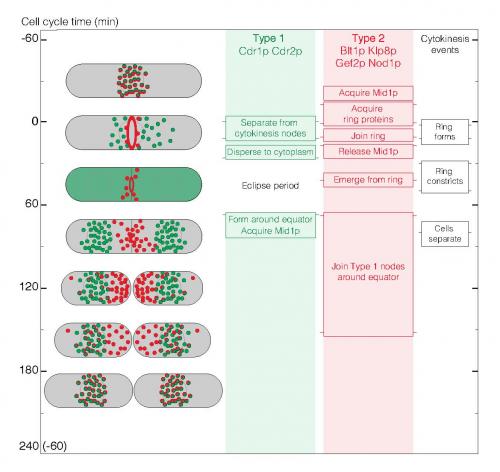
Image above: Summary of the ~5 hour node cycle in fission yeast cells. The left column shows cartoons of cells through the cell cycle with Type 1 nodes in green, Type 2 nodes in red and colocalized Types 1 and 2 nodes in red/green. The other columns indicate the times of events during the cell cycle. The light green and red background shading shows when node markers are organized in structures. Horizontal lines on the side of each box mark the average time of each event. Pairs of horizontal lines represent the beginning and end of events spread over time.
