Project Overview
The project area we describe below lies at the interface of wet lab experimental work and mathematical modeling.
The physical properties of bacteria are still poorly understood, despite their importance on cellular processes. Generally, the cytoplasm, where most cellular processes occur, is assumed to behave as a simple (viscous) liquid. Collaborative work between the Jacobs-Wagner, Dufresne and O’Hern labs has suggested otherwise.
We have found that the bacterial cytoplasm behaves like a colloidal suspension on the cusp of a glass transition. Glassy materials are characterized by dynamic arrest and can be fluidized by relatively subtle perturbations, a role that cellular metabolism appears to fulfill in the bacterial cytoplasm.
These findings are opening novel areas of research as they imply many interesting biological and physical questions. From the physical perspective, colloidal glasses are fascinating materials, but are typically studied using monodisperse particles. In contrast, the bacterial cytoplasm is a highly polydisperse material (its components’ sizes range over 3 orders of magnitude), and we expect continued work on the bacterial cytoplasm to inform us about glassy dynamics in complex environments. From the biological perspective, we expect glassy properties to have wide-ranging implications for bacterial physiology.
For example, metabolic dormancy is a prevalent bacterial state in nature (including clinical settings), and we will explore how bacteria may have exploited glassy properties over the course of evolution to resist stressful environments such as starvation and antibiotic treatment. Specific projects will include experiments, modeling, or both according to the student’s interest.
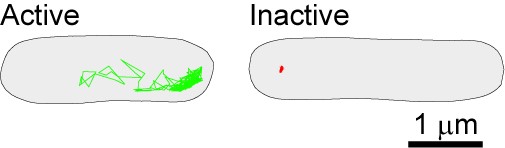
Image above: The bacterial cytoplasm displays dramatically different physical properties depending on the metabolic state of the cell, as shown by probe confinement in inactive cells (red trajectory in image above) and probe exploration of the cytoplasmic space (green trajectory). In this experiment, the probes (GFP-µNS particles) were tracked at a 1.3 sec frame rate for 650 sec.
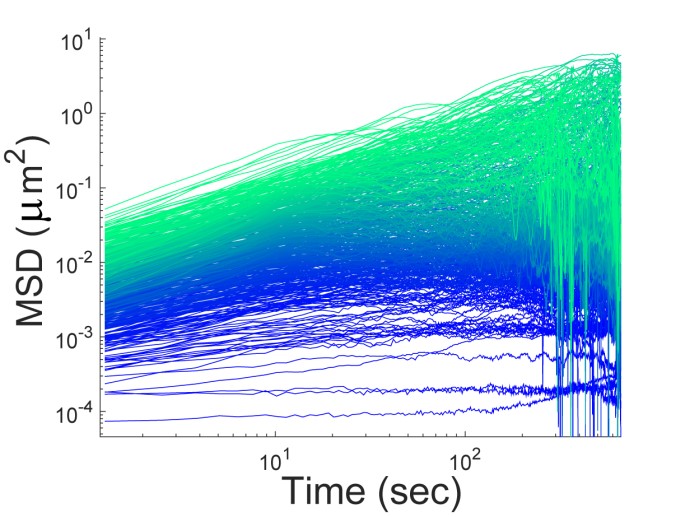
Image above: Single-particle tracking in genetically-identical E. coli cells growing under the same conditions reveals a surprisingly large heterogeneity in particle motion, as shown in the single trajectory mean square displacements (MSDs) above. We are interested in understanding the origin of this heterogeneity and its role in biology (e.g., phenotypic cell-to-cell variability).
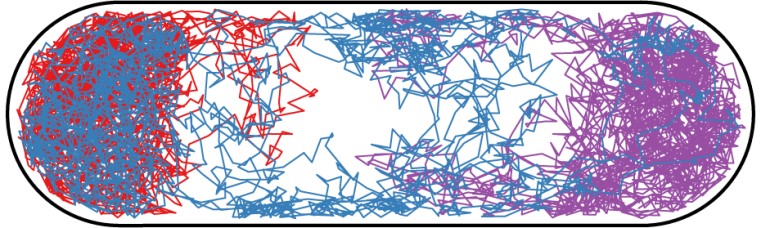
Image above: We are developing models of the bacterial cytoplasm. The figure above shows three simulated trajectories of a particle diffusing through the cytoplasm of E. coli. The cell was modeled in 3-dimensions with a hard membrane and a non-penetrable inner spherocylindrical region that represents the DNA region (nucleoid) of the cell.
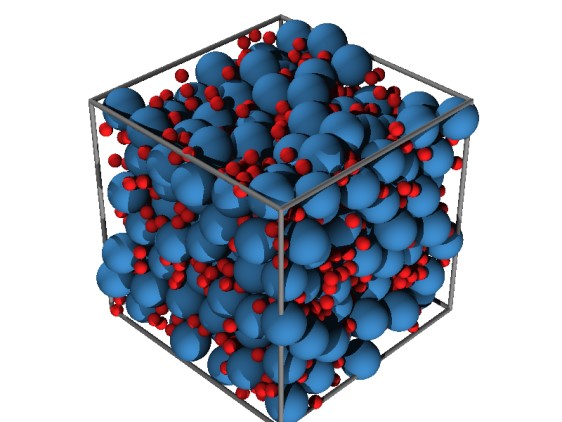
Image above: We also use modeling to understand the effect of crowding and polydispersity in the cytoplasm. The image above shows an example of packing of 1,000 spherical particles of two different sizes.
